Unmonitored reservoirs of influenza A virus in pigs could pose potential epidemic early warning problems

Research led by a team at Duke-National University of Singapore Medical School, Singapore, has found alarmingly active swine flu reservoirs in South East Asia. Their study traced the evolutionary origin of these viruses and identified migration patterns.
In a paper, “The genomic landscape of swine influenza A viruses in Southeast Asia,” published in PNAS, the team details the co-circulation of multiple swine influenza lineages in Cambodian pig populations, including diverse reassortant viruses and their genetic origins.
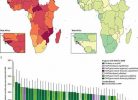
From March 2020 to July 2022, the researchers conducted swine influenza surveillance in 18 pig slaughterhouses in Cambodia. They collected 4,089 nasal swabs from pigs in different districts of four provinces. Among these, 72 pigs (1.8%) tested positive for influenza A virus, with the highest positivity rate (4.5%) found in Kandal province compared to other provinces (ranging from 0.2% to 1.8%).
They analyzed complete or partial swine influenza genomes from 45 nasal swab samples. H1 and H3 HA subtypes and N1 and N2 NA subtypes were identified, with co-infections observed. The predominant subtype was H1N1, present in 82.2% of the sequenced samples.
Phylogenetic analysis revealed diverse lineages of H1 in circulation within Cambodia, with most H1 and N1 sequences being derivatives of the human H1N1/pdm09 lineage. H3 influenza subtypes were found in 22.2% of the samples, often co-infecting with H1N1/pdm09 viruses.
While the current study sampled pigs in Cambodia, the problem has been reported across Southeast Asia and is related to virus reservoirs elsewhere in the world. For example, European avian-like H1N2 was detected. The findings also suggest that a North American swine N2 segment may have been introduced into Asia over 15 years ago.
From an epidemiologist’s perspective, this introduction should have been noticed over a decade ago in order to be able to react to an outbreak or jump to humans. This requires systematic surveillance of swine influenza viruses, which is not taking place in Southeast Asia and has limited compliance in many nations.
From a pig farmer’s perspective, there can be nothing to indicate a potential problem, nothing wrong with the health of the pigs or outward signs that they are harboring multiple forms of influenza A, and it does not affect pork products or food health.
Monitoring viruses is expensive and requires a different set of skills than available on a typical hog farm. The implications of an unmonitored domestic pig population acting as a reservoir for multiple forms of influenza to cohabitate and potentially recombine into a deadly pandemic could be far more costly.
Before SARS-CoV-2
Before COVID-19 became a household name, the epidemic most on the minds of epidemiologists was the influenza A virus. The influenza A virus is found in some wild animals, and it tends to remain there without affecting large populations until it finds a domestic animal host. From there, the virus comes in closer contact with other animals and people and can begin to adapt to other hosts. A variant of the influenza A virus is also responsible for bird flu.
The 1918 “Spanish flu,” which likely jumped hosts on a chicken farm in the United States but was first noticed by soldiers stationed in Spain, was an avian-sourced H1N1 influenza A virus. The outbreak infected ~500 million people globally, or ~33% of the world’s population, from 1918 to 1919, resulting in ~50 million deaths.
In 2009, a swine flu (also H1N1) pandemic infected millions of people worldwide, and thanks to the quick work of scientists around the world, people can be protected from H1N1 with an annual flu shot, orange juice and antiviral medications. Even with vaccination efforts, the influenza A virus is estimated to be responsible for ~450,000 deaths worldwide each year.
More information:
Michael A. Zeller et al, The genomic landscape of swine influenza A viruses in Southeast Asia, Proceedings of the National Academy of Sciences (2023). DOI: 10.1073/pnas.2301926120
Journal information:
Proceedings of the National Academy of Sciences
Source: Read Full Article