SARS-CoV-2 Brazilian variant may potentially escape T cell-mediated host immune responses
A team of scientists from Brazil recently explored the abilities of the UK, South African, and Brazilian variants of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) to evade host CD8+ T cell responses induced by natural infection or vaccination. The findings reveal reduced antigenic coverage for the P.2 Brazilian lineage, indicating its ability to evade SARS-CoV-2-induced T cell responses. The study is currently available on the bioRxiv* preprint server.
.jpg)
Background
Since the emergence of coronavirus disease 2019 (COVID-19) pandemic, efforts are continuously being made to sequence the SARS-CoV-2 genome and examine potential viral mutations. Although the mutation rate of SARS-CoV-2 is considerably lower than other RNA viruses, some potential spike mutations have been identified in recently emerged variants of SARS-CoV-2 (the UK, South African, and Brazilian variants) that can significantly increase viral infectivity. A growing pool of evidence also indicates that some of these variants can evade host cell humoral immune responses induced by natural infection or vaccination. In the context of COVID-19, it has been observed that, unlike relatively short-lived anti-SARS-CoV-2 antibody responses, T cell-mediated adaptive immune responses provide robust and long-lasting protection against SARS-CoV-2. This suggests the need for developing new COVID-19 vaccines with specific peptides that can induce appropriate T cell responses against SARS-CoV-2.
In the current study, the scientists have evaluated the impact of new spike- and nucleocapsid-specific SARS-CoV-2 mutations on T cell-mediated host immune responses. Specifically, they analyzed whether these mutations influence viral peptide coverage among the human population; human leukocyte antigen (HLA)-mediated presentation of viral peptides to T cells; and overall T cell responses.
Study design
Specifically, the scientists studied the impacts of spike/nucleocapsid mutations (26 single nucleotide substitution mutations and 2 deletion mutations) presents in four potential SARS-CoV-2 variants (UK variant: B.1.1.7 Lineage; South African variant: B.1.351 Lineage; and two Brazilian variants: Lineages P.1 and P.2) using the sequence of original Wuhan strain as a wild-type reference sequence.
Initially, they conducted binding prediction experiments to identify viral peptides that bind either strongly (strong binders) or weakly (weak binders) to a set of class I HLA molecules obtained from 29 countries. For peptide antigenicity prediction, they selected a set of viral peptides that were distinct between each viral variant and the wildtype reference strain. To estimate population coverage for each viral variant, they used class I HLA: peptide pairs obtained from the wildtype SARS-CoV-2 and its variants.
Important observations
By conducting the binding prediction analysis, the scientists identified a set of 230 unique viral peptides that generated 6320 HLA-I: peptide combinations. Importantly, they observed that the spike N501Y mutation was associated with 10 times higher production of viral peptides than the wildtype sequence. Moreover, all substitution mutations present in the nucleocapsid encoding region generated more peptides than the wildtype sequence. These observations indicate that the mutations in the spike protein and nucleocapsid of newly emerged SARS-CoV-2 variants may actually increase the number of viral peptides presented to the HLA-I molecules, which in turn can trigger the CD8+ T cell-mediated cytotoxic responses and terminate the viral replication/propagation.
Furthermore, the scientists determined the affinities of wildtype- and variant-derived peptides for the HLA-I molecules. They observed that a set of six peptides derived from the T205I, D80A, H655Y, N501Y, P26S, and T716I substitution mutations exhibited significantly increased affinity for the HLA-I molecules. Importantly, the spike mutations present in the UK variant generated peptides with significantly increased affinity, whereas a significantly reduced affinity was observed for peptides derived from one of the Brazilian variants (Lineage P.2).
By considering all single nucleotide substitution mutations present in each SARS-CoV-2 variant, they observed an increased antigenicity for the South African and P.1 Brazilian variants. Interestingly, they observed that some of the mutation-derived peptides could be more antigenic but less presented by the HLA-I molecule. Similarly, some peptides could be less antigenic but better presented by the HLA-I molecule. These findings indicate that mutations present in the immunodominant region of the wildtype sequence can actually facilitate the viral evasion of host T cell responses.
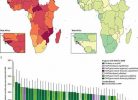
Furthermore, they estimated the global antigenic coverage for the four SARS-CoV-2 variants and observed a lower antigenic coverage for the P.2 Brazilian variant and higher coverage for the South African and P.1 Brazilian variants.
Study significance
The study reveals that viral peptides derived from the P.2 Brazilian variant of SARS-CoV-2 are poorly presented by the HLA-I molecules, indicating its potency to escape CD8+ T cell-mediated host immune responses induced either by natural infection or vaccination. Moreover, the study indicates that peptides derived from the spike N501Y and nucleocapsid T205I mutations may have higher affinity for HLA-I molecules compared to the wildtype SARS-CoV-2 sequence.
*Important Notice
bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
- Pretti MAM. 2021. New SARS-CoV-2 lineages could evade CD8+ T-cells response. BioRxiv. doi: https://doi.org/10.1101/2021.03.09.434584, https://www.biorxiv.org/content/10.1101/2021.03.09.434584v1
Posted in: Medical Science News | Medical Research News | Disease/Infection News | Healthcare News
Tags: Antibody, Antigen, Cell, Coronavirus, Coronavirus Disease COVID-19, Genome, Human Leukocyte Antigen, Leukocyte, Molecule, Mutation, Nucleotide, Pandemic, Peptides, Propagation, Protein, Respiratory, RNA, SARS, SARS-CoV-2, Severe Acute Respiratory, Severe Acute Respiratory Syndrome, Spike Protein, Syndrome

Written by
Dr. Sanchari Sinha Dutta
Dr. Sanchari Sinha Dutta is a science communicator who believes in spreading the power of science in every corner of the world. She has a Bachelor of Science (B.Sc.) degree and a Master's of Science (M.Sc.) in biology and human physiology. Following her Master's degree, Sanchari went on to study a Ph.D. in human physiology. She has authored more than 10 original research articles, all of which have been published in world renowned international journals.
Source: Read Full Article