Identifying SARS-CoV-2 epitopes targeted by current vaccination could help improve future vaccines
Researchers in the United States have identified antibody-inducing epitopes of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) in individuals vaccinated against coronavirus disease 2019 (COVID-19) that could be used for future diagnostics, therapeutics, and vaccine design.
Using an approach called reverse vaccinology, the team identified distinct amino acid motifs and epitopes of the SARS-CoV-2 spike protein that elicit specific immunoglobulin IgG antibody responses that have previously been associated with neutralization of the virus.
The spike protein is the main structure SARS-CoV-2 uses to infect cells and the primary target of neutralizing antibodies following vaccination or natural infection.
-2.jpg)
The team – from Yale University School of Medicine in New Haven, Connecticut and Serimmune, Inc. in Goleta, California – identified dominant linear epitopes in subunit 1 (S1) and subunit 2 (S2) of the spike protein that have previously been associated with SARS-CoV-2 neutralization in vitro.
Spike S1 contains the receptor-binding domain (RBD) that attaches to the host cell receptor angiotensin-converting enzyme 2 (ACE2) as the initial stage of the infection process, while S2 contains the machinery that enables the fusion of the viral envelope to the cell membrane.
Adam Wisnewski and colleagues say that the epitopes identified here should be considered in the context of future diagnostics, therapeutic design and vaccine development.
A pre-print version of the research paper is available on the medRxiv* server, while the article undergoes peer review.
The epitopes that induce SARS-CoV-2-specific IgG remain incompletely characterized
The first vaccines to have been authorized for emergency use during the COVID-19 pandemic elicit antibodies that exhibit significant SARS-CoV-2 neutralizing capacity and protect against severe disease.
However, the spike protein epitopes that induce SARS-CoV-2-specific IgG in vaccine recipients and could form the basis of future “epitope vaccines” remain incompletely characterized, says Wisnewski and colleagues.
Reverse vaccinology is an evolving approach for improving vaccine effectiveness and minimizing adverse responses by limiting immunizations to critical epitopes,” adds the team.
Although knowledge about the immunogenic SARS-CoV-2 spike epitopes in vaccinated individuals is limited, immunodominant epitopes in naturally infected patients have been identified based on recognition by serum IgG.
The spike protein contains conformational epitopes in the RBD and dominant linear epitopes in the C-terminal domains of S1 and S2 and the fusion peptide region of S2.
Importantly, dominant linear epitopes and specific conformational epitopes have been shown to mediate viral neutralization in vitro,” say the researchers.
What did the current study involve?
The team used a reverse vaccinology approach to identify spike protein epitopes that are targeted following immunization with either the Pfizer-BioNTech BNT162b2 vaccine or Moderna’s mRNA-1273 vaccine.
Using a technique called Serum Epitope Repertoire Analysis (SERA), the researchers mapped the immunogenic amino acid motifs and linear epitopes of the SARS-CoV-2 spike protein that elicit IgG in paired pre- and post-immunization samples taken from 20 adults without prior COVID-19 who received the Pfizer-BioNTech vaccine.
SERA’s unbiased, whole proteome approach to epitope discovery is ideal for mapping the targets of humoral responses to vaccination,” says Wisnewski and colleagues.
The analysis of pre- versus post-vaccination samples was conducted to help ensure immune specificity resulted from vaccination rather than pre-existing cross-reactivity.
The analysis was also applied to eight individuals without prior COVID-19 who received the Moderna vaccine and five individuals with prior COVID-19 who were immunized with either the Pfizer-BioNTech (n=2) or the Moderna (n=3) vaccine.
What did the study find?
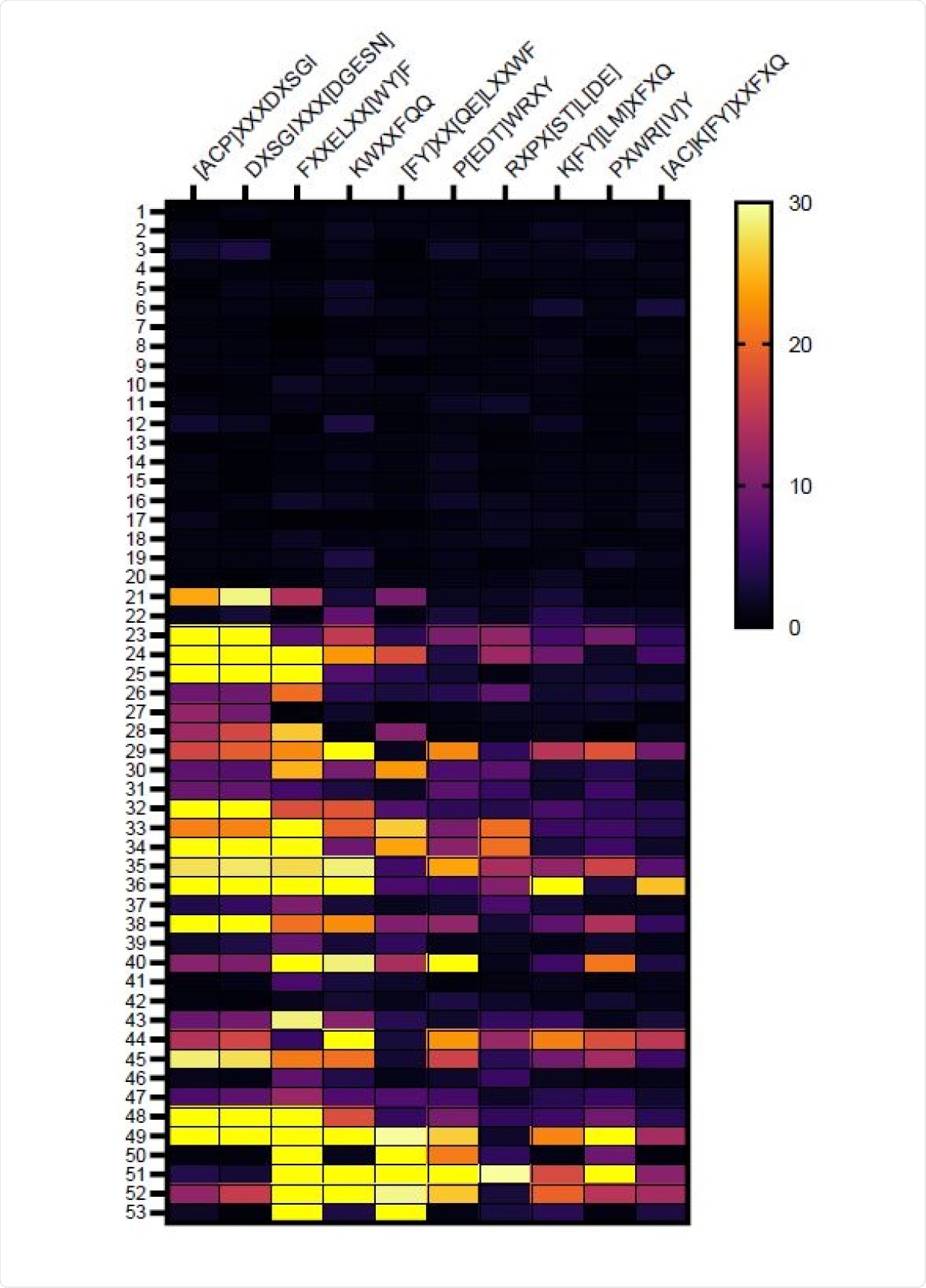
The team identified several distinct amino acid motifs recognized by vaccine-induced IgG that may reflect 3-dimensional conformations.
Dominant linear epitopes were identified in the C-terminal domains of the S1 and S2 subunits (amino acids 558-569, 627-638, and 1148-1159) that have previously been associated with SARS-CoV-2 neutralization.
In summary, we identified epitopes of COVID-19 mRNA vaccines that trigger specific IgG responses previously associated with SARS-CoV-2 neutralizing activity,” says Wisnewski and colleagues.
These epitopes may form the basis of future diagnostics, therapeutics, and focused vaccine development, concludes the team.
*Important Notice
medRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
- Wisnewski A, et al. Immunogenic Amino Acid Motifs and Linear Epitopes of COVID-19 mRNA Vaccines. medRxiv, 2021. doi: https://doi.org/10.1101/2021.05.25.21257427, https://www.medrxiv.org/content/10.1101/2021.05.25.21257427v1
Posted in: Medical Science News | Medical Research News | Miscellaneous News | Disease/Infection News | Healthcare News
Tags: ACE2, Amino Acid, Angiotensin, Angiotensin-Converting Enzyme 2, Antibodies, Antibody, Cell, Cell Membrane, Coronavirus, Coronavirus Disease COVID-19, Diagnostics, Enzyme, Immunization, Immunoglobulin, in vitro, Medicine, Membrane, Pandemic, Protein, Proteome, Receptor, Research, Respiratory, SARS, SARS-CoV-2, Severe Acute Respiratory, Severe Acute Respiratory Syndrome, Spike Protein, Syndrome, Therapeutics, Vaccine, Virus

Written by
Sally Robertson
Sally first developed an interest in medical communications when she took on the role of Journal Development Editor for BioMed Central (BMC), after having graduated with a degree in biomedical science from Greenwich University.
Source: Read Full Article