An Overview of Protein Quantification Methods
Elucidating information about proteins present in biological samples is vital for scientists working in the field of life sciences. Whilst qualitative data can tell us which proteins are present in a given sample, knowing the number of proteins in a sample can provide invaluable data and results. This article will provide a brief overview of commonly used protein quantification methods and why they are so useful.
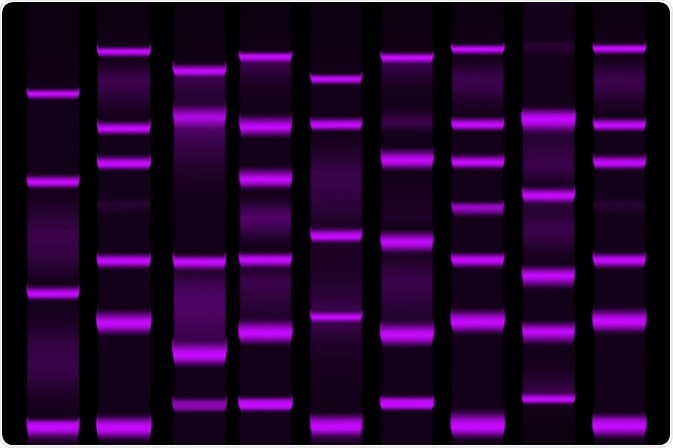 Image Credit: extender_01 / Shutterstock.com
Image Credit: extender_01 / Shutterstock.com
Protein Quantification – A Brief Overview
In any study which seeks to isolate, purify, characterize, and analyze protein samples, protein quantification should always be part of the researcher’s toolkit. The techniques used are identical to qualitative methods, with quantification as an added dimension.
By analyzing the amount of protein in a target of interest, methods provide information about physiological differences between two samples compared to just identifying the proteins present. One example would be the difference between healthy and infected cells.
The term for the field in analytical chemistry which studies proteins quantitatively is Quantitative proteomics. There are many different methods available to researchers to study the makeup of proteins that have been developed over the past few decades. Common assays used in protein quantification methods include the Bradford, Folin-Lowry, and Bicinchoninic Acid (BCA) assays, each with its own advantages and disadvantages.
Quantitative proteomics has diverse applications in biomedical fields including drug and biomarker discovery. The techniques in the field have started to supersede ELISA and western blot methods.
Methodologies
There are several methods available for studies. These include mass spectrometry, 2-dimensional gel electrophoresis, spectrophotometry, DIGE, QDB, and others. Methods make use of fluorescent dyes to aid in identification.
Each technique has advantages and disadvantages compared to other methods, due to the dynamic and complex nature of protein samples in living tissue. Techniques require the use of highly specialized equipment.
Two-dimensional gel electrophoresis
2-DE provides information on quantity, charge, and mass. Limitations become apparent in this method once it is used to analyze protein samples larger than 150kDa or smaller than 5kDa as well as proteins with low solubility. 2-DE also requires MS techniques for downstream protein identification.
Difference gel electrophoresis (DIGE) is a variant of 2D-E with higher sensitivity. Up to three protein samples can be labeled with fluorescent cyanine-based dyes including Cy3 and Cy2 covalently bound to proteins prior to 2-DE analysis. This brings an extra dimension of statistical reliability and confidence. The dyes are mass and charge-matched but, crucially, have unique fluorescent properties. They migrate simultaneously but produce distinct excitation and emission spectra.
Mass Spectrometry
Mass spectrometry methods are one of the main methods for quantitative protein analysis. Quantitative MS has high sensitivity but only provides limited information about intact proteins. Quantitative MS can either populations of cells (bulk analysis) or individual cells (single-cell analysis.)
Early spectrometric approaches applied isotope-coded affinity tags. Developed in the 1990s, this laid the groundwork and foundation for quantitative proteomics. The tags use two reagents (with heavy and light isotopes respectively) and a biotin affinity tag. One development in the field using these tags in conjunction with mass spectrometry was the labeling of whole S. cerevisiae cells.
Since the use of these tags, methods have expanded to include isobaric mass tags, stable isotope labels, tandem mass tags, metal-coded tags, stable-isotope labeling with amino acids in cell culture (SILAC), and many others.
One major disadvantage in using mass spectrometry methods is that the intensity of a peak in a mass spectrum is not a good indicator of amounts of analyte. It is not inherently quantitative – differences in ionization effectiveness and/or detectability of the numerous peptides in a sample. However, differences in peak intensity between the same analyte in different samples can accurately reflect abundance.
Mass spectrometry is therefore commonly used in conjunction with several other methods to produce reliable and accurate quantification data.
Quantitative Dot Blot
A recent technique that has been developed is quantitative dot blot. The main advantage of QDB is that it can measure both absolute and relative quantities of individual proteins in a high-throughput manner. It can significantly reduce the time needed to quantify proteins compared to other methods such as 2D-E due to not requiring complex blotting procedures. However, QDB does not provide information on protein size.
QDB is a variant of the popular and simple western blot method.
Label-free Quantification Methods
Label-free quantification methods are a relatively cheap and simple set of methods. In label-free quantification, samples are separately analyzed and their spectra are compared to determine relative peptide abundance between them. Whilst it is cheap and simple, it is the least accurate quantification paradigm (although reliable when subjected to heavy statistical validation.)
The two different methods of label-free quantification are AUC (area under the curve) and spectral counting. Accuracy can be improved in label-free methods by the use of high-resolution spectrometers and specialized computer software.
In Conclusion
Quantitative protein analysis and the field of quantitative proteomics is a constantly evolving scientific discipline. As drug development and biomedical science advances further into the 21st century, the use of analytical chemistry techniques such as this will no doubt gain precedence, helping scientists peer ever-deeper into the dynamics of cellular environments.
Sources
Yu, G et al. (2020) Developing a routine lab test for absolute quantification of HER2 in FFPE breast cancer tissues using Quantitative Dot Blot (QDB) method Nature Scientific Reports 10 Article No. 12502 [Accessed Online 5th October 2020] https://www.nature.com/articles/s41598-020-69471-4
Rupprecht, K.R. et al. (2010) Development of a dot-blot assay for screening monoclonal antibodies to low-molecular-mass drugs Anal. Biochem. Vol. 407 Issue 2, Pp. 160-164 [Accessed Online 5th October 2020] www.sciencedirect.com/science/article/abs/pii/S0003269710005014
Pan, S. et al. (2009) Mass Spectrometry Based Targeted Protein Quantification: Methods and Applications J. Proteome Res. Vol. 8, Issue 2 Pp. 787-797[Accessed Online 5th October 2020] https://pubs.acs.org/doi/abs/10.1021/pr800538n
Further Reading
- All Protein Content
- Protein Production: Initiation, Elongation and Termination
- Protein Folding
- Amino Acids and Protein Sequences
- Protein Complex Analysis
Last Updated: Dec 17, 2020

Written by
Reginald Davey
Reg Davey is a freelance copywriter and editor based in Nottingham in the United Kingdom. Writing for News Medical represents the coming together of various interests and fields he has been interested and involved in over the years, including Microbiology, Biomedical Sciences, and Environmental Science.
Source: Read Full Article